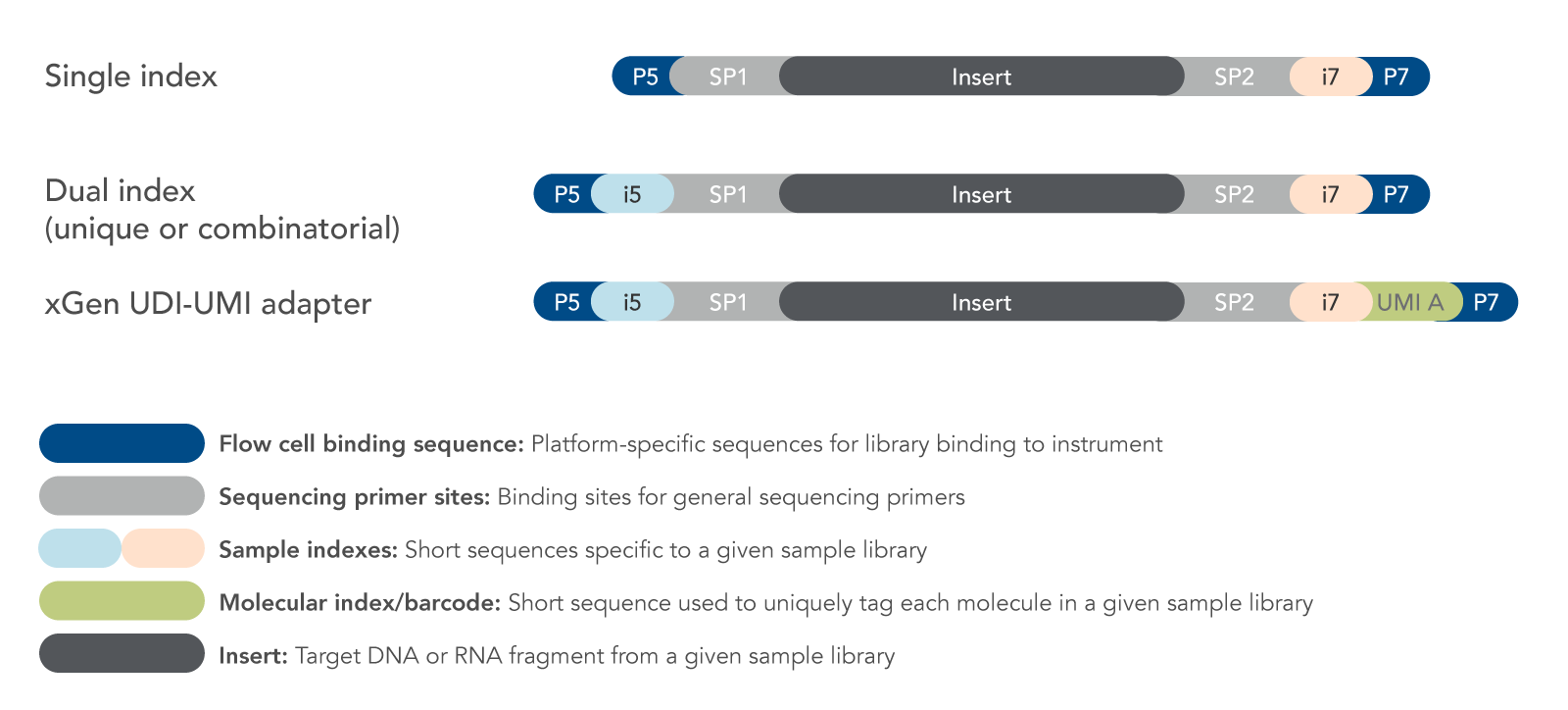
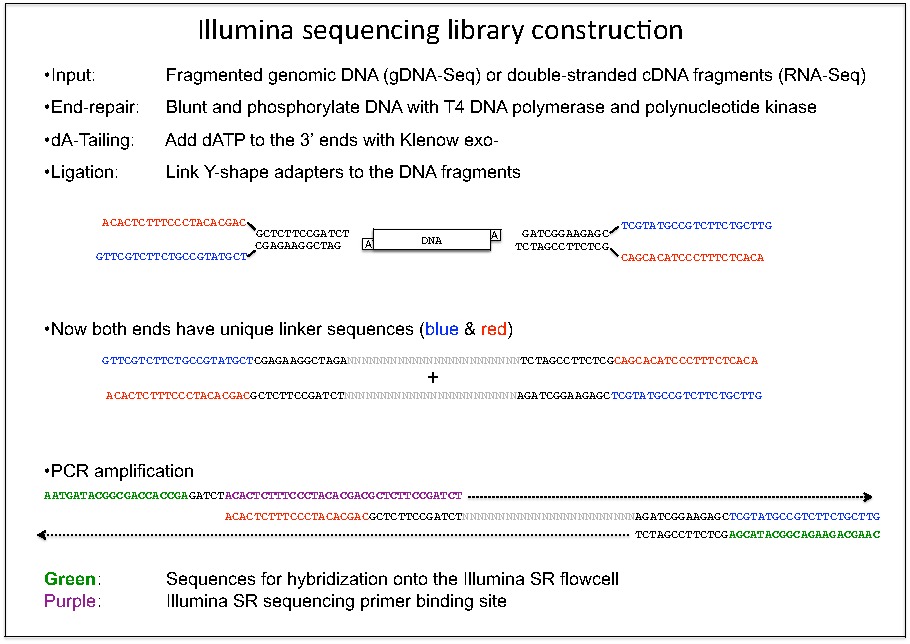
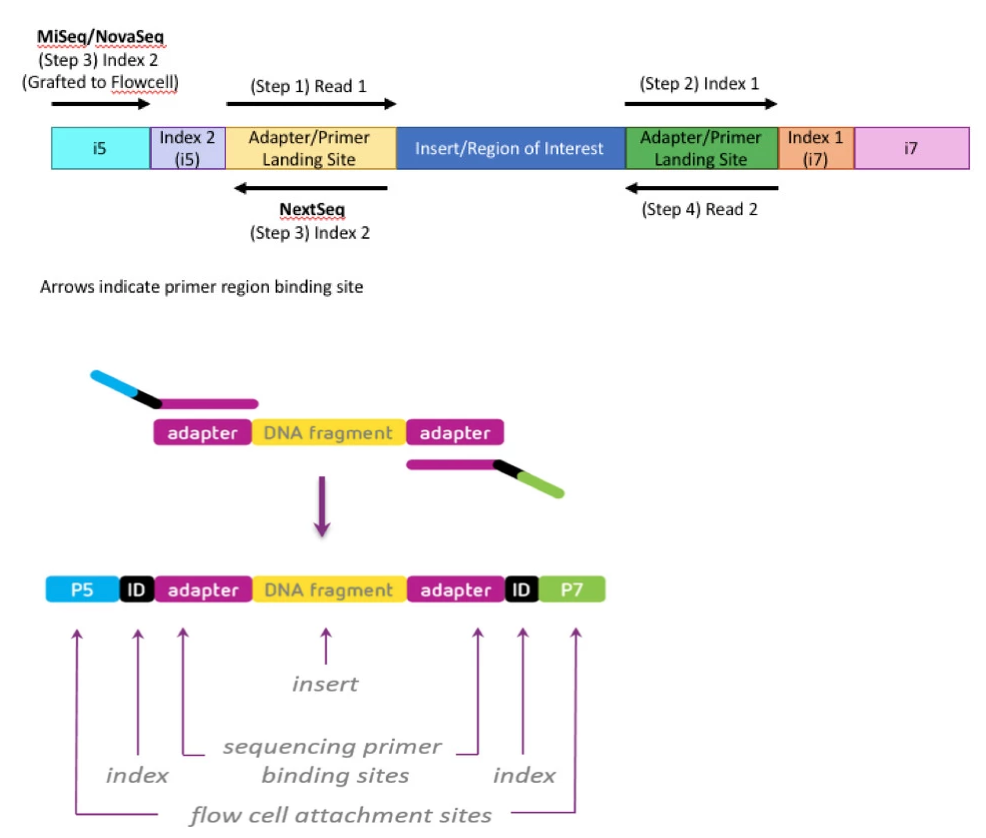
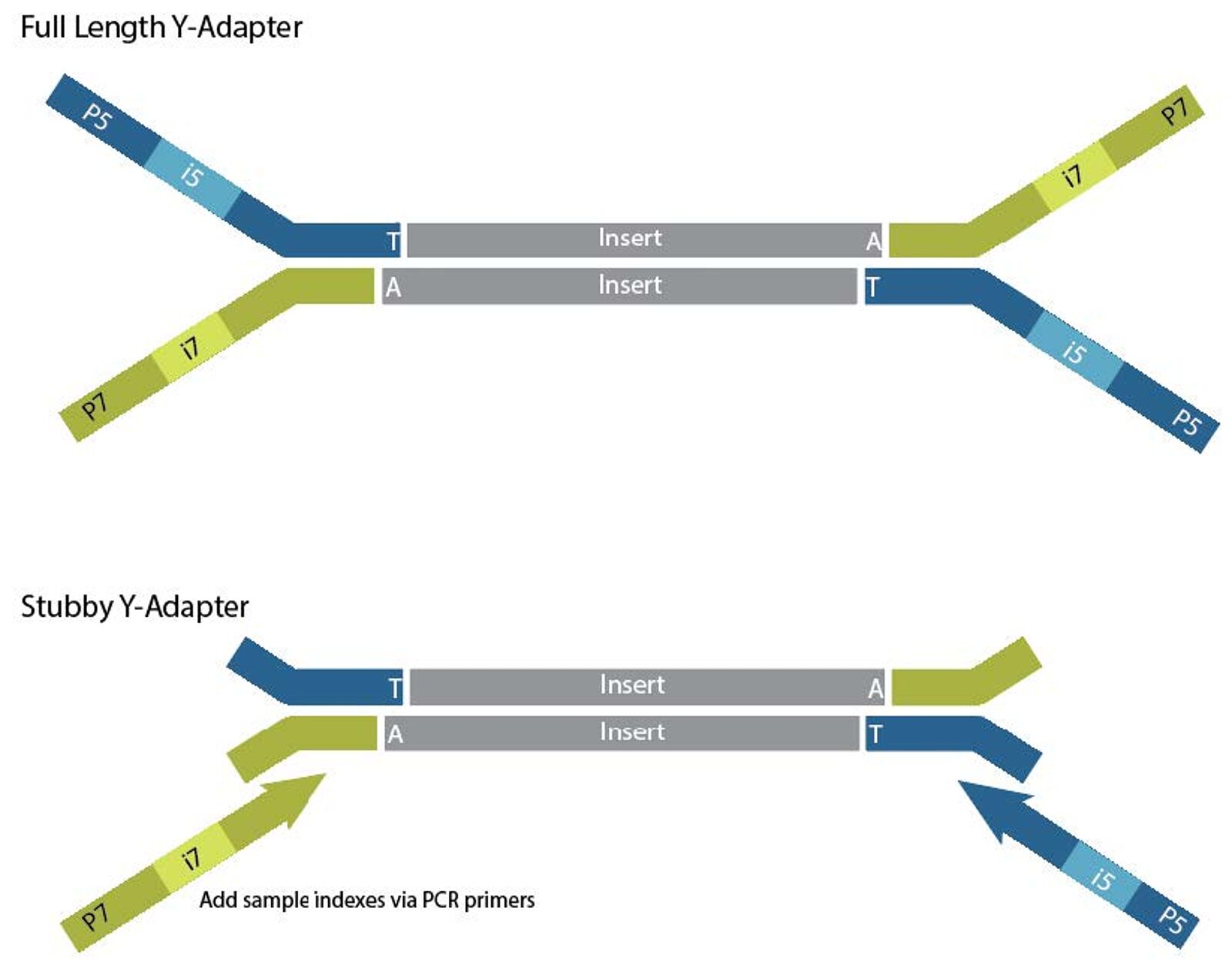
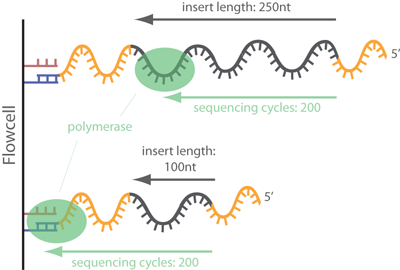
Adapter trimming Why are adapter sequences trimmed from only the 3' ends of reads Adapter trimming: Why are adapter sequences trimmed from only the 3' ends of reads - Illumina Knowledge

NGS Adapters in Ligation-Based Library Prep Workflows | Biocompare: The Buyer's Guide for Life Scientists
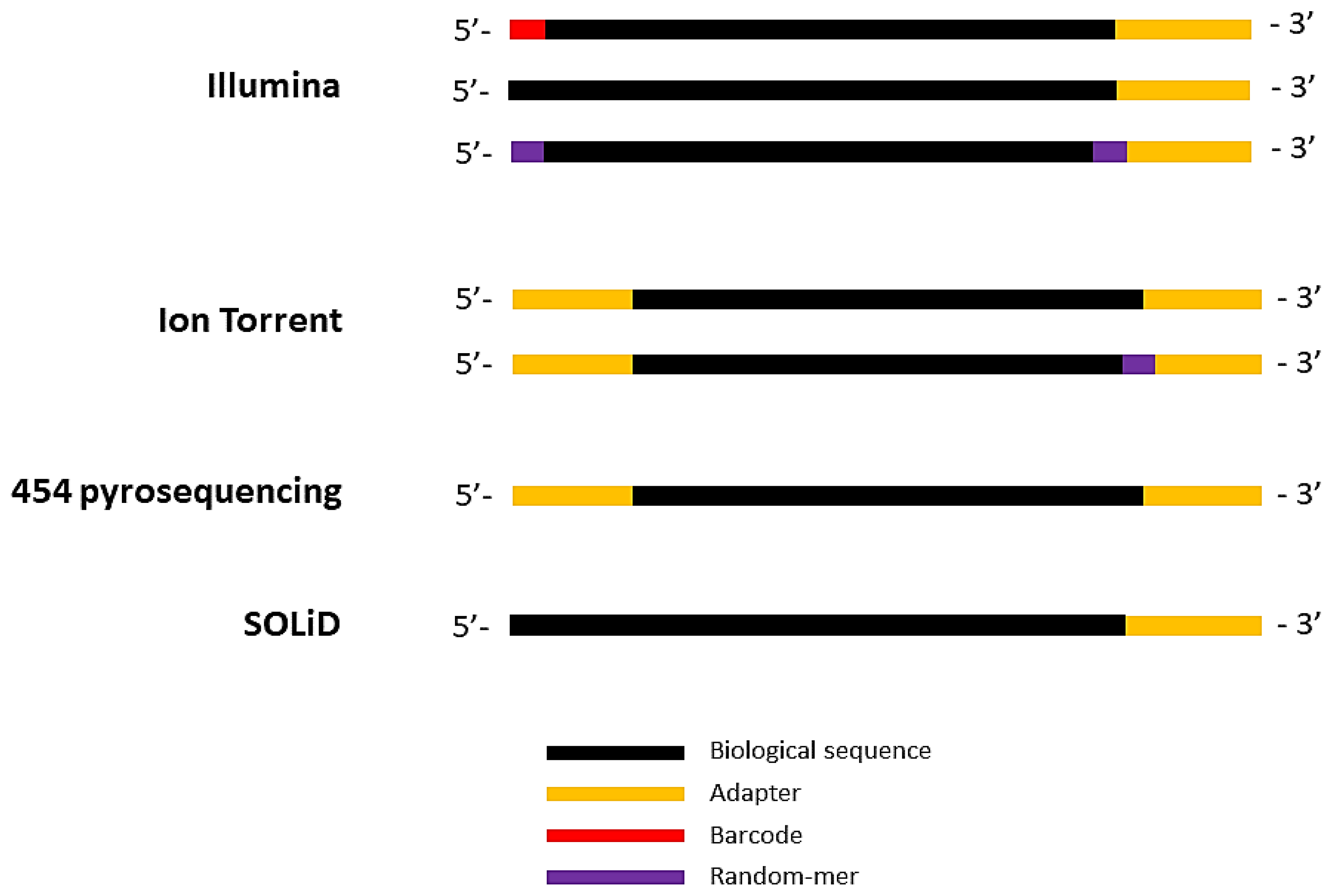
Biomolecules | Free Full-Text | High-Throughput Identification of Adapters in Single-Read Sequencing Data

Ultrahigh-Throughput Multiplexing and Sequencing of >500-Base-Pair Amplicon Regions on the Illumina HiSeq 2500 Platform | mSystems
High-Throughput, Amplicon-Based Sequencing of the CREBBP Gene as a Tool to Develop a Universal Platform-Independent Assay | PLOS ONE



![Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ] Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7786/1/fig-1-full.png)